About
CRISPResso is a software pipeline designed to enable rapid and intuitive interpretation of genome editing experiments. A limited web implementation is available at: http://crispresso2.pinellolab.org/ or http://crispresso.com.
Briefly, CRISPResso:
- Aligns sequencing reads to a reference sequence
- Quantifies insertions, mutations and deletions to determine whether a read is modified or unmodified by genome editing
- Summarizes editing results in intuitive plots and datasets
Tools
CRISPResso is a suite of complementary tools:
- CRISPResso - for analyzing and interpreting single experimental conditions on a single amplicon
- CRISPRessoBatch - for analyzing and comparing multiple experimental conditions at the same site
- CRISPRessoPooled - for analyzing multiple amplicons from a pooled amplicon sequencing experiment
- CRISPRessoWGS - for analyzing specific sites in whole-genome sequencing samples
- CRISPRessoCompare - for comparing editing between two samples (e.g., treated vs control)
- CRISPRessoAggregate - for aggregating results from previously-run CRISPResso analyses
How can you use CRISPResso?
CRISPResso can be used to analyze genome editing outcomes using cleaving nucleases (e.g. Cas9 or Cpf1) or noncleaving nucleases (e.g. base editors). The following operations can be automatically performed:
- Filtering of low-quality reads
- Adapter trimming
- Alignment of reads to one or multiple reference sequences (in the case of multiple alleles)
- Quantification of HDR and NHEJ outcomes (if the HDR sequence is provided)
- Quantification frameshift/inframe mutations and identification affected splice sites (if an exon sequence is provided)
- Visualization of the indel distribution and position (for cleaving nucleases)
- Visualization of distribution and position of substitutions (for base editors)
- Visualization of alleles and their frequencies
CRISPResso processing
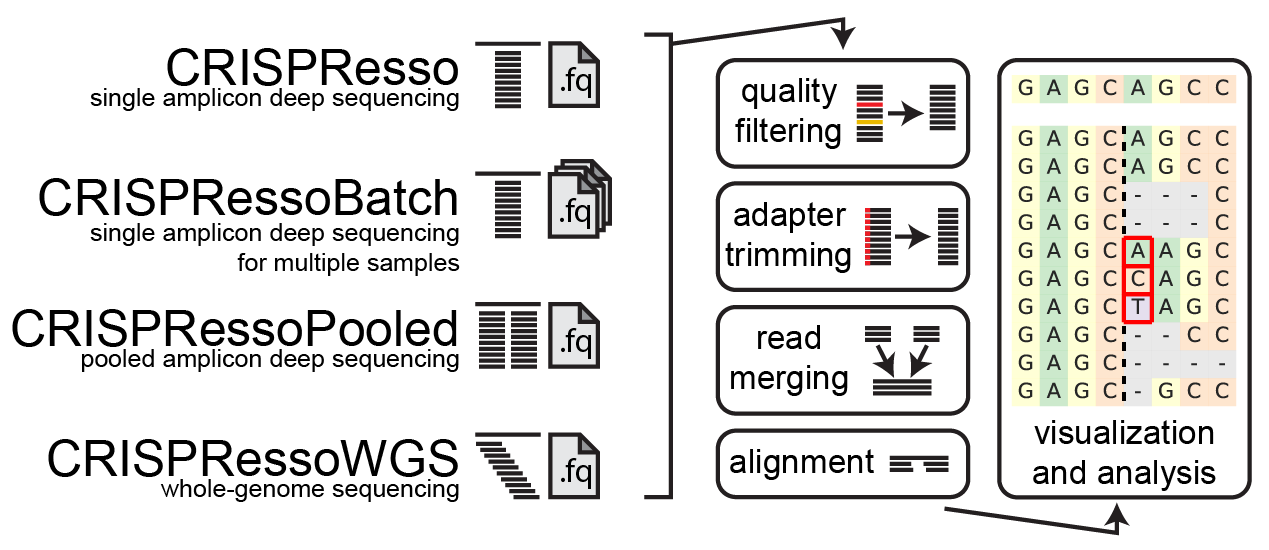
Quality filtering
Input reads are first filtered based on the quality score (phred33) in order to remove potentially false positive indels. The filtering based on the phred33 quality score can be modulated by adjusting the optimal parameters (see additional notes below).
Adapter trimming
Next, adapters are trimmed from the reads. If no adapter are present, select 'No Trimming' under the 'Trimming adapter' heading in the optional parameters. If reads contain adapter sequences that need to be trimmed, select the adapters used for trimming under the ‘Trimming adapter’ heading in the optional parameters. Possible adapters include Nextera PE, TruSeq3 PE, TruSeq3 SE, TruSeq2 PE, and TruSeq2 SE. The adapters are trimmed from the reads using fastp.
Read merging
If paired-end reads are provided, reads are merged using fastp. This produces a single read for alignment to the amplicon sequence, and reduces sequencing errors that may be present at the end of sequencing reads.
Alignment
The preprocessed reads are then aligned to the reference sequence with a global sequence alignment algorithm that takes into account our biological knowledge of nuclease function. If multiple alleles are present at the editing site, each allele can be passed to CRISPResso and sequenced reads will be assigned to the reference sequence or origin.
Visualization and analysis
Finally, after analyzing the aligned reads, a set of informative graphs are generated, allowing for the quantification and visualization of the position and type of outcomes within the amplicon sequence.
How is CRISPResso2 different from CRISPResso?
CRISPResso2 introduces four key innovations for the analysis of genome editing data:
- Comprehensive analysis of sequencing data from base editors. We have added additional analysis and visualization capabilities especially for experiments using base editors.
- Allele specific quantification of heterozygous references. If the targeted editing region has more than one allele, reads arising from each allele can be deconvoluted.
- A novel biologically-informed alignment algorithm. This algorithm incorporates knowledge about the mutations produced by gene editing tools to create more biologically-likely alignments.
- Ultra-fast processing time.
Installation
CRISPResso can be installed using the conda package manager Bioconda, or it can be run using the Docker containerization system.
Bioconda
To install CRISPResso using Bioconda, download and install Anaconda Python, following the instructions at: https://docs.anaconda.com/free/anaconda/install/.
Open a terminal and type:
conda config --add channels defaults
conda config --add channels bioconda
conda config --add channels conda-forge
To install CRISPResso into the current conda environment, type:
conda install crispresso2
Alternatively, to create a new environment named crispresso2_env with CRISPResso, type:
conda create -n crispresso2_env -c bioconda crispresso2
Activate your conda environment:
conda activate crispresso2_env
Verify that CRISPResso is installed using the command:
CRISPResso -h
Bioconda for Apple Silicon
If you would like to install CRISPResso using bioconda on a Mac with Apple silicon (aren't sure?), then there is a slight change you need to make. First, ensure that you have Rosetta installed. Next, you must tell bioconda to install the Intel versions of the packages. If you would like to do this system wide, which we recommend, run the command:
conda config --add subdirs osx-64
Then you can proceed with the installation instructions above.
If you would like to use the Intel versions in a single environment, then run:
CONDA_SUBDIR=osx-64 conda create -n crispresso2_env -c bioconda crispresso2
If you choose to use the CONDA_SUBDIR=osx-64 method, note that if you install additional packages into the environment you will need to add the CONDA_SUBDIR=osx-64 to the beginning of each command. Alternatively, you could set this environment variable in your shell, but we recommend to use the conda config --add subdirs osx-64 method because it is less error prone.
Docker
CRISPResso can be used via the Docker containerization system. This system allows CRISPResso to run on your system without configuring and installing additional packages. To run CRISPResso, first download and install docker: https://docs.docker.com/engine/installation/.
Next, Docker must be configured to access your hard drive and to run with sufficient memory. These parameters can be found in the Docker settings menu. To allow Docker to access your hard drive, select 'Shared Drives' and make sure your drive name is selected. To adjust the memory allocation, select the 'Advanced' tab and allocate at least 4G of memory.
To run CRISPResso, make sure Docker is running, then open a command prompt (Mac) or Powershell (Windows). Change directories to the location where your data is, and run the following command:
docker run -v ${PWD}:/DATA -w /DATA -i pinellolab/crispresso2 CRISPResso -h
The first time you run this command, it will download the Docker image. The -v parameter mounts the current directory to be accessible by CRISPResso, and the -w parameter sets the CRISPResso working directory. As long as you are running the command from the directory containing your data, you should not change the Docker -v or -w parameters.
Additional parameters for CRISPResso as described below can be added to this command. For example,
docker run -v ${PWD}:/DATA -w /DATA -i pinellolab/crispresso2 CRISPResso -r1 sample.fastq.gz -a ATTAACCAAG
Troubleshooting
Please check that your input file(s) are in FASTQ format (compressed fastq.gz also accepted).
If you get an empty report, please double check that your amplicon sequence is correct and in the correct orientation. It can be helpful to inspect the first few lines of your FASTQ file - the start of the amplicon sequence should match the start of your sequences. If not, check to see if the files are trimmed (see point below).
It is important to determine whether your reads are trimmed or not. CRISPResso2 assumes that the reads ARE ALREADY TRIMMED! If reads are not already trimmed, select the adapters used for trimming under the ‘Trimming Adapter’ heading under the ‘Optional Parameters’. This is FUNDAMENTAL to CRISPResso analysis. Failure to trim adaptors may result in false positives. This will result in a report where you will observe an unrealistic 100% modified alleles and a sharp peak at the edges of the reference amplicon in figure 4.
The quality filter assumes that your reads uses the Phred33 scale, and it should be adjusted for each user’s specific application. A reasonable value for this parameter is 30.
If your amplicon sequence is longer than your sequenced read length, the R1 and R2 reads should overlap by at least 10bp. For example, if you sequence using 150bp reads, the maximum amplicon length should be 290 bp.
Especially in repetitive regions, multiple alignments may have the best score. If you want to investigate alternate best-scoring alignments, you can view all alignments using this tool: http://rna.informatik.uni-freiburg.de/Teaching/index.jsp?toolName=Gotoh. As input, sequences from the 'Alleles_frequency_table.txt' can be used. Specifically, for a given row, the value in the 'Aligned_Sequence' should be entered into the 'Sequence a' box after removing any dashes, and the value in the 'Reference_Sequence' should be entered into the 'Sequence b' box after removing any dashes. The alternate alignments can be selected in the 'Results' panel in the Output section.
Changelog
Unreleased
ADDED
-
Add an amino acid nucleotide quilt plot by @mbowcut2 in #552
-
Add
scripts/reconstituteReads.pyto generate FASTQ from CRISPResso2 output by @kclem ina800762andcd79dcc -
Add an UpSet plot to represent bystander edits for Base Editing analyses by @mbowcut2 in #554
-
Allow for messages to be served via CRISPResso reports by @Colelyman in #583
-
Add a plot that shows the distribution of homology scores for reads by @mbowcut2 in #600
FIXED
-
Fix parsing the
CRISPResso2_info.jsonin CRISPRessoPooled by @kclem in #558 -
Forced cloned
include_idxsto benp.arrays by @kclem inda4badb -
Fix the link to the CRISPResso cup in reports (so that SSL works correctly) by @Colelyman in #571
-
Fix the quantification of deletions at the second position of the sequence by @Colelyman in #574
-
Fix an issue with unaligned reads not being reported correctly when writing BAM output by @trevormartinj7 in #578
-
Fix an issue where quantification window coordinates we not being correctly inferred by @Colelyman in #598
- This issue is present when there is a single quantifcation window coordinate provided and multiple amplicons. What happens is CRISPResso aligns the second amplicon to the first and then infers what the quantification window coordinates should be based on the alignment. A regression was introduced where the inference of the quantification window coordinates for the second amplicon was no longer correct. This change fixes the regression and brings the behavior back to match that of v2.2.9.
- If you don't set quantification window coordinates and don't use multiple amplicons, there is no need for this fix and therefore no change in behavior.
-
Fix a
SyntaxWarningfor an unescaped sequence in a matplotlib function by @Colelyman in #600 -
Fix a bug during
--bam_outputwhen there is an unaligned read, the remainder of the reads will not bu output by @Colelyman in #602
CHANGED
- Update the base Docker image to
mambaorg/micromamba:2.3.3and remove dependency on Anacondadefaultschannel by @Colelyman in #575
REMOVED
v2.3.3 - Activity Fulton - 07/01/2025
ADDED
-
Asymmetrical Allele Plots by @mbowcut2 and @Colelyman in #527
-
Native CRISPResso Paired End Read Merging by @Colelyman and @Snicker7 in #537
-
Support for amplicon names with emojis 🎉🤩 and other non-standard characters by @kclem in
1455609 -
Multiplexing for CRISPRessoPooled subruns in
1455609
FIXED
- Fix setting of 99%ile in negative direction for deletions plot in
90ac42a
CHANGED
-
Make fig_filename_root default to None, in which case the figure is shown interactively (e.g. in a jupyter notebook) in
c2a10c4 -
Don't rerun if --no_rerun is set but --verbosity has changed in
6562a08
REMOVED
- Remove warning for zipping nonexistant files in
3784ea5
v2.3.2 - Junction Salt - 01/16/2025
ADDED
-
New parameters,
--flexiguide_gap_open_penaltyand--flexiguide_gap_extend_penalty, to customize flexiguide alignment in #491 -
New parameter
--halt_on_plot_failso that errors and exceptions in plots don't fail silently in #494 -
New parameter
--samtools_exclude_flagto customize the filtering of reads in #503 -
New documentation website at <docs.crispresso.com>.
-
d3 plot enhancements by @trevormartinj7 in #459
-
Add flexiguide alignment parameters by @Colelyman in #491
-
Add pyproject.toml and support numpy v2 by @Snicker7 in #496
-
Add customizable samtools exclude flag by @Colelyman in #503
-
Add support for octal and comma separated samtools exclude flags (#113) by @kclem in #507
FIXED
-
Fix typo and move flexiguide to debug (#77) by @Colelyman in #438
-
Matplotlib Compatibility Fix by @mbowcut2 and @Snicker7 in #464
-
Fix CRISPRessoAggregate bug and other improvements (#95) by @Colelyman in #470
-
Fix missing substitution in name of WGS, Compare and Meta reports by @Colelyman in #498
-
Fix
get_n_fastqfunction by @trevormartinj7 in #508
CHANGED
-
Improvement of processing speed by
-
Progress percentages are displayed in the CLI output.
-
Prefix the release Docker tag with a
vby @Colelyman in #434 -
Pin versions of numpy and matplotlib in CI environment by @Snicker7 in #452
-
Implement new pooled mixed-mode default behavior by @mbowcut2 in #454
-
Update README by @Snicker7, @mbowcut2, @trevormartinj7, @Colelyman and @kclem in #456
-
Cache conda packages in GIthub Actions by @Colelyman in #466
-
Replace zcat by @Colelyman in #468
-
Cache read merging step in CRISPRessoPooled on no_rerun by @kclem in #467
-
Display percentages in the CLI output by @Colelyman in #473
-
No processor pool when running in single thread by @Snicker7 in #474
-
Round percentage complete in CLI and add initial 0% complete by @Colelyman in #477
-
Reduce memory usage for allele plots by @Colelyman in #478
-
Sync reports by @Colelyman in #479
-
Read Alignment Parallelization (#98) by @trevormartinj7 in #480
-
Add
all_deletion_coordinatesto be returned byfind_indels_substitutions_legacyfunction by @Colelyman in #486 -
Update jinja_partials and bring Reports into sync by @Colelyman in #500
-
Update detailed alleles table help option by @Colelyman in #513
v2.3.1 - Screen King - 05/13/2024
FIXED
-
Extract
jinja_partialsand fix CRISPRessoPooled fastp errors by @Colelyman and @trevormartinj7 in #425 -
Fix batch mode pandas warning. (#70) by @mbowcut2 and @Colelyman in #429
-
Fix issues with
file_prefixby @Colelyman and @Snicker7 in #430 -
Fix plots and improve plot error handling by @Snicker7 and @mbowcut2 in #431
CHANGED
-
Cole/refactor jinja undefined (#66) by @Colelyman and @Skicker7 in #421
-
Update README by @Colelyman in #424
-
Bump version to 2.3.1 and change default CRISPRessoPooled behavior to change in 2.3.2 by @Colelyman in #428
-
Showing sgRNA sequences on hover in CRISPRessoPro by @Colelyman in #432
REMOVED
- Remove extra imports from CRISPRessoCore by @Colelyman in #422
v2.3.0 - Targeting Minato - 04/10/2024
ADDED
-
Guardrails (checking experimental conditions and raising warnings) by @Snicker7
FIXED
-
Fix samtools piping by @Colelyman in #325
-
Fix interleaved fastq input in CRISPRessoPooled and suppress CRISPRessoWGS params by @Colelyman in #392
-
Fix #367, reads only align to prime edited amplicon, not to reference by @mbowcut2 in #393
-
Fix the assignment of multiple quantification window coordinates by @Snicker7 in #403
-
Fix Jinja2 undefined variables by @Colelyman in #417
CHANGED
-
Flash and Trimmomatic are replaced with Fastp by @trevormartinj7, @Snicker7, and @Colelyman
-
Failed runs are displayed with the error by @trevormartinj7
-
Replace link to CRISPResso schematic with raw URL in README by @Colelyman in #329
-
Run unit tests via Github Actions and fix matplotlib character issue by @Snicker7 and @mbowcut2 in #386
-
Remove future Pandas warnings and sort CRISPRessoCompare tables by @mbowcut2 and @Snicker7 in #389
-
Move read filtering to after merging in CRISPResso by @Colelyman in #397
-
Decrease Docker image size and fix PE naming and parameter behavior by @Colelyman, @Snicker7 and @mbowcut2 in #404
v2.2.14 - Specific São Paulo - 08/10/2023
FIXED
v2.2.13 - With Montgomery - 07/28/2023
ADDED
- Add verbosity argument to CRISPRessoAggregate (#18) fixes #306 by @Colelyman in #307
FIXED
-
Parallel plotting fix by @Colelyman and @kclem in
546446eand #286 -
Fix multiprocessing lambda pickling by @Colelyman in #311
CHANGED
-
Don't start pool when only using single thread by @Colelyman in #302
-
Raise exceptions from within futures in plot_pool in
a439f09 -
Enable CRISPRessoPooled multiprocessing when os allows multi-thread file append in
ebb016d -
Allow multiple overlapping sgRNA matches in reference (previous behavior was to only search for non-overlapping sgRNA sites in the reference sequence in
32e1e97 -
Assert correct input fastq file format in
7248ba8 -
Update plotCustomAllelePlot.py script for #292 by @kclem in #293
-
Clarify CRISPRessoWGS intended use by @Colelyman in #303
-
Case-insensitive headers accepted in CRISPRessoPooled
e577318 -
Allow dashes in filenames in
712eb2a -
Sort pandas dataframes by # of reads and sequences so that the order is consistent for testing by @Snicker7 and @Colelyman in #316
-
Update
base_editorparameters in README and add Plot Harness by @Colelyman in #301
v2.2.12 - Protospace Utah - 02/01/2023
ADDED
-
Add deprecation notice in #260
-
Add snippet about installing CRISPResso2 via bioconda on Apple silicon in #274
FIXED
CHANGED
- Status Updates + Pooled Mixed Mode Update in #279
v2.2.11 - Of Weber - 10/11/2022
FIXED
-
Fix batch quilt plot name by @Colelyman in #249
-
Fix typo of CRISPResssoPlot when plotting nucleotide quilt by @Colelyman in #250
v2.2.10 - Overhangs Alameda - 09/15/2022
ADDED
-
Add
--zip_outputparameter to produce a zipped file report by @Colelyman and @Snicker7 inc80f828 -
Autodetect reference amplicons from interleaved fastq input
FIXED
-
Fix bug when comparing two samples with the same name in #228
-
Fix bug when name is provided instead of amplicon_name in pooled input file in #229
-
Fix for aggregate plots in Batch mode in #237
-
Fix loading of crispressoInfo from WGS and pooled in
49740ba
CHANGED
v2.2.9 - Long Surrey - 06/23/2022
ADDED
- fastq_to_bam implementation in #219\
- If the parameter --bam_output is provided, CRISPResso alignments will be written to a file called 'CRISPResso_output.bam' with the alignments in bam format. If the
bowtie2_indexis provided, alignments will be reported in reference to that genome. If thebowtie2_indexis not provided, alignments will be reported in reference to a custom reference created by the amplicon sequence(s) and written to the file 'CRISPResso_output.fa'.\ - This enables the viewing of CRISPResso alignments in other browsers (e.g., IGV). If no
bowtie2_indexis provided, the reference genome should be set to the produced 'CRISPResso_output.fa' file, and then the alignment bam can be loaded into IGV.
- If the parameter --bam_output is provided, CRISPResso alignments will be written to a file called 'CRISPResso_output.bam' with the alignments in bam format. If the
FIXED
- Don't run global frameshift plot when there are no modified reads by @Colelyman in #226
CHANGED
v2.2.8 - Welcome to High Waikato - 05/13/2022
ADDED
-
Interactive plotly summary plots in CRISPRessoAggregate and CRISPRessoBatch for visualizing and comparisons
-
CRISPRessoPooled enhancement that allows the amplicons file to have a header and additional columns to be provided
-
CRISPRessoCompare generates a report of the number of significant reads at each base
CHANGED
-
Minor bug fixes for plotCustomAllelePlot.py to work with Python3 by @dharjanto in #212
-
Large aggregation by @Colelyman in #192
v2.2.7 - Literature and Los Angeles - 02/11/2022
ADDED
- Adds features for providing aligned bams as input to CRISPRessoPooled and for a faster demultiplexing when amplicons and genome are provided. The added parameters are:
--aligned_pooled_bam: Path to aligned input for CRISPRessoPooled processing. If this parameter is specified, the alignments in the given bam will be used to demultiplex reads. If this parameter is not set (default), input reads provided by--fastq_r1(and optionally--fastq_r2) will be aligned to the reference genome using bowtie2. If the input bam is given, the corresponding reference fasta must also be given to extract reference genomic sequences via the parameter--bowtie2_index. Note that the aligned reads are paired-end seqenced, they should already be merged into 1 read (e.g. via Flash) before alignment.--demultiplex_only_at_amplicons: If set, and an amplicon file (--amplicons_file) and reference sequence (--bowtie2_index) are provided, reads overlapping alignment positions of amplicons will be demultiplexed and assigned to that amplicon. If this flag is not set, the entire genome will be demultiplexed and reads with the same start and stop coordinates as an amplicon will be assigned to that amplicon.
FIXED
-
Fix int bug for CRISPRessoPooled n_reads (
ef15cae)
CHANGED
-
Improve performance by removing regex from indel location analysis by @Colelyman #182
-
Fastq output produced by
--fastq_outputnow includes the inserted bases. Previously, a string like "DEL= INS=78(1) SUB= " would indicate a 1bp insertion at site 78. This update outputs strings like "DEL= INS=78(1+G) SUB= " with the insertion described as a plus character followed by the inserted bases. (2f84dd0) -
Allow mixed-case prime-editing input (
e999079)
v2.2.6 - Basepairing Bern - 10/21/2021
ADDED
-
Add param --plot_center to allow custom plots centered at a given point in plotCustomAllelePlot script ecf23ef
-
Add unit tests 3e6c281
FIXED
-
Fix allele plotting error for plot 5 referring to uninitialized y_max variable 53197e6
-
Fix unicode errors for bam read/write 8196b6a
CHANGED
-
All sub-CRISPResso runs are run with 1 process in Batch, WGS, Pooled, etc. Because we added multiprocessing capabilities to CRISPResso (the plotting part) we thought it would be slick for CRISPRessoPooled to run sqrt(n_processes) CRISPResso processes with sqrt(n_processes) processes each. Unfortunately, sqrt(n) isn't a really useful number for the number of processes people usually run (e.g. 2 or 3 or even 8), and a lot of the CRISPResso processing isn't enabled to take advantage of multiprocessing (e.g. the alignment step), so processes were being wasted. So we reverted back to having n_processes CRISPResso processes, each with 1 process. a923a7c
-
Convert columns in nucleotide count and modification tables to numeric for PE analysis cabebbe
-
Make loggers module-specific so matplotlib debug doesn't get spewn out in the CRISPResso log c2bdd96
REMOVED
- Remove version checks for numpy and seaborn 90b43ea
v2.2.5 - Immunity from Bonneville - 09/22/2021
FIXED
- Fixes bug when sequencing reads are much longer than the given reference sequence.
v2.2.4 - Mutant Maricopa - 09/09/2021
ADDED
- This release adds an additional parameter --assign_ambiguous_alignements_to_first_allele. For ambiguous alignments, setting this flag will force them to be assigned to the first (as provided by the references -a first and then -e second) amplicon. Thus, no reads will be discarded as 'ambiguous' and all reads will be counted once in the analysis.
CHANGED
- Batch summaries are produced for amplicons present in only one sample.
v2.2.3 - Collateral Cardston - 08/30/2021
FIXED
- Fixes database_id bug
v2.2.2 - Large Honolulu - 08/20/2021
FIXED
- For some reason some of the previous commits resolving problems with filterFastqs weren't picked up in v2.2.1. So I'm hoping they'll be included here.
v2.2.1 - Sequence Length Salt Lake - 08/20/2021
FIXED
- More unicode bug fixes for filtering fastqs
CHANGED
- CRISPRessBatch now outputs summary of splicing/frameshift mutation status
v2.2.0 - Matches Sanpete - 08/13/2021
CHANGED
-
Python 3 release
-
Incorporates updates and changes from python2 up to this point
-
Adds multiprocessing to CRISPRessoBase to parallelize image generation to speed up results
- CRISPRessoBatch, CRISPRessoPooled, and CRISPRessoWGS allocate processes to sub-CRISPResso commands so that sqrt(n_processes) sub-commands are run, each with sqrt(n_processes) unless plotting is turned off (via --suppress_report or --suppress_plots in which case n_processes are run, each with 1 process.
-
The crispresso_info dictionary containing run information is saved as json so it can be read across versions of python and by other programs (e.g. R). The dict structure of the object has also been changed to be more navigable and hierarchical.
-
Because of changes to crispresso_info, this version of CRISPResso will not be able to finish incomplete runs (e.g. checkpointing of CRISPRessoPooled) that were started by previous python2 versions of CRISPResso, or aggregate or access information from previous runs (e.g. using CRISPRessoAggregate, CRISPRessoCompare, or the custom python plotting scripts in scripts). However, even if we were to have stuck with pickle format for crispresso_info, the python2 and python3 versions were incompatible anyway. So we figured this was a good time to move toward a better format.
-
v2.1.3 - Lentiviral Cache - 06/29/2021
FIXED
- Fixes bug for CRISPRessoPooled analyzes of many amplicons where samtools sort writes status updates that can't be parsed as reads.
v2.1.2 - Single Guide Washington - 06/23/2021
ADDED
- Addition of a script for custom allele plotting 947fbab
CHANGED
- Updates to CRISPRessoPooledWGSCompare, used for comparing multiple amplicons in CRISPRessoWGS or CRISPRessoPooled experiments 48d6c87
- CRISPRessoPooledWGSCompare now produces html report linking to sub-CRISPRessoCompare reports
v2.1.1 - Nicking Hancock - 05/22/2021
ADDED
- This release incorporates changes to make bowtie2 alignment in CRISPRessoPooled more permissive 4dc9e7, and remove duplicate rows in the Alleles_frequency_table.txt due to reads being in the forward or reverse direction 0e08cd0.
CHANGED
- When given a genome file, CRISPRessoPooled aligns reads to the genome using the Bowtie2 aligner. The legacy parameters were somewhat strict. The new parameters reflect the 'default_min_aln_score' parameter in allowing for substantially more indels and mismatches than previous.
- The parameter --use_legacy_bowtie2_options_string has been added to use the legacy settings. Otherwise, the bowtie2 alignment settings will be calculated as follows:
- --end-to-end - no clipping, match bonus -ma is set to 0
- -N 0 number of mismatches allowed in seed alignment
- --np 0 where read (or ref have ambiguous character (N)) penalty is 0
- --mp 3,2 mismatch penalty - set max mismatch to -3 to coincide with the gap extension penalty (2 is the default min mismatch penalty)
- --score-min L,-5,-3*(1-H) For a given homology score, we allow up to (1-H) mismatches (-3) or gap extensions (-3) and one gap open (-5). This score translates to -5 + -3(1-H)L where L is the sequence length
v2.1.0 - Knockout Lake - 03/23/2021
CHANGED
-
Starting in version 2.1.0, insertion quantification has been changed to only include insertions completely contained by the quantification window. To use the legacy quantification method (i.e. include insertions directly adjacent to the quantification window) please use the parameter --use_legacy_insertion_quantification
-
In Prime Editing mode pegRNA spacer sequences given in the incorrect orientation are no longer tolerated
-
HDR: Ambiguous alignments don't contribute to the plot 4g (except when --expand_ambiguous_alignments is provided) --fastq_output now also writes alignment scores and alignments for every read
v2.0.45 - 12/30/2020
ADDED
- CRISPRessoAggregate can be used to aggregate multiple completed CRISPResso runs.
v2.0.44 - 11/17/2020
CHANGED
- Improvements in inferring quantification windows across amplicons/alleles.
v2.0.43 - 11/07/2020
ADDED
-
Add ticks to appropriate plots
-
New parameter --plot_histogram_outlier to plot 100% of data
FIXED
-
Update the function of histograms
-
By default 99% of data is shown in plots, now 100% of data is written to data files.
v2.0.42 - 09/30/2020
FIXED
- Fixed % character in CRISPRessoPooled arg string
v2.0.41 - 09/30/2020
ADDED
- Added --fastq-out parameter to report the CRISPResso analysis separately for each read. Note that this should be used with caution. I'm still trying to figure out what information should be reported for each read, and what format it should be in. Open to feedback on this issue!
FIXED
- WGS parallelization mode bug fixed
CHANGED
- WGS and Pooled summary figures scale height based on the number of entries so that they are legible in html reports.
v2.0.40 - 07/09/2020
CHANGED
- Prime editing updates - scaffold parameter is now called --prime_editing_pegRNA_scaffold_seq. Guide names with spaces produce file names with hyphens instead of spaces
v2.0.39 - 07/07/2020
ADDED
- Batch mode supports bam and multiple quantification windows
v2.0.38 - 07/01/2020
ADDED
-
Add new paramter --annotate_wildtype_allele to annotate wildtype alleles on the allele plots
-
Input can now be read from bam using the parameter --bam_input and (optionally) --bam_chr_loc to use the reads in the bam at this location as input.
-
An output bam is produced with an additional soace-separated field prefixed by c2 (e.g. c2:Z:ALN=Inferred CLASS=Inferred_MODIFIED MODS=D47;I0;S0 DEL=56(47) INS= SUB= ALN_REF=TTGGCGGATGTTCCAATCAGTACGCAGAGAGTCGCCGTCTCCAAGGTGAAAGCGGAAGTAGGGCCTTCGCGCACCTCATGGAATCCCTTCTGCAGCACCTGGATCGCTTTTCCGAGCTTCTGGCGGTCTCAAGCACTACCTACGTCAGCACCTGGGACCCCGCCACCGTGCGCCGGGCCTTGCAGTGGGCGCGCTACCTGCGCCACATCCATCGGCGCTTTGGTCGGCATGGCCCCATTCGCACGGCTCT----------------------------------------------- ALN_SEQ=ACACCGGATGTTCCAATCAGTACGCAGAGAGTCGCCGTCTCCAAGGTGAAAGCGGA-----------------------------------------------TCGCTTTTCCGAGCTTCTGGCGGTCTCAAGCACTACCTACGTCAGCACCTGGGACCCCGCCACCGTGCGCCGGGCCTTGCAGTGGGCGCGCTACCTGCGCCACATCCATCGGCGCTTTGGTCGGCATGGCCCCATTCGCACGGCTCTGGAGCGGCGGCTGCACAACCAGTGGAGGCAAGAGGGCGGCTTTGGGC). Note that the alignment details (location, cigar string, etc) are not modified.. this may be done in the future). Bam file input cannot be trimmed or pre-processed with quality filtering.
CHANGED
-
Prime editing scaffold incorporation is now more accurate (looks for the scaffold sequence at the expected position directly after the extension sequence). A plot showing the number of bases matching the scaffold, as well as insertions after the extension sequence, and a data file with these numbers is produced. Added parameter --prime_editing_pegRNA_scaffold_min_match_length to define the minimum length required to classify a read as 'Scaffold-incorporated'
-
Renamed split_paired_end parameter to --split_interleaved_input for interleaved input
-
Auto mode now considers 5000 reads to detect amplicon sequences
-
Update output when reporting missing files -- only lists first 15 files in the current directory and directory of input parameter
-
--reference https instead of http
v2.0.37 - 05/09/2020
ADDED
-
Max processors can be used in WGS and Pooled modes by setting -p max
-
Prime editing analysis can be performed by specifying the parameters: --prime_editing_pegRNA_spacer_seq --prime_editing_pegRNA_extension_seq and optionally --prime_editing_pegRNA_extension_quantification_window_size --prime_editing_pegRNA_scaffold_sequence --prime_editing_nicking_guide_seq with a summary shown in the report
-
Extended read analysis data available with --write_detailed_allele_table flag
CHANGED
-
CRISPRessoPooled demultiplexing is performed in parallel and with reduced filesystem demand
-
N's don't count as substitutions
-
Nucleotide plots are shaded when the nucleotide matches the reference sequence
-
sgRNA improvements: sgRNA annotations are plotted on multiple lines if they overlap sgRNAs can have their own cut site and quantification window size
v2.0.34 - 04/06/2020
ADDED
- Pooled Set flag to skip reporting problematic regions
v2.0.33 - 04/03/2020
CHANGED
-
Plotting computation window is shaded
-
Parallelization and checkpointing of CRISPRessoWGS and Pooled
-
Increase of alignment efficiency of CRISPRessoPooled amplicons in genome +amplicons mode
v2.0.32 - 02/25/2020
CHANGED
- Plotting updates, dsODN detection, and general improvements and bug fixes.
v2.0.31 - 09/26/2019
ADDED
- Add custom post-processing plot functions for allele tables
FIXED
- Fix CRISPRessoPooled handling of chromosomes with underscores
CHANGED
- Update dependency requirements
v2.0.30 - 07/02/2019
ADDED
- Add nucleotide summary for batch mode
FIXED
- Fix bug for reporting amplicons with no reads
CHANGED
- Case-insensitive checking for guides
v2.0.29 - 05/30/2019
CHANGED
-
By default, the html report is created on the outside of the output folder, so if the output is: CRISPResso_on_SAMPLE/ the html report will be at CRISPResso_on_SAMPLE.html
-
This functionality can be reverted to place the report inside of the output folder using the parameter --place_report_in_output_folder which will place the html report at: CRISPResso_on_SAMPLE/CRISPResso2_report.html
v2.0.28 - 05/24/2019
ADDED
-
Standardize file names
-
Add CRISPREssoCompare output html
CHANGED
-
CRISPRessoBatch guide-specific output are plotted as separate plots
-
Standardize window definitions (plot window and quantification window specify the distance from the cut site to the edge of the window, so the entire window is 2*plot window)
v2.0.27 - 04/05/2019
Added
-
Add reports for pooled and WGS
-
Add Batch pickle info
CHANGED
-
More precise plotting of cleavage cut site and quantification window
-
Bioconda updates
v2.0.26 - 03/06/2019
Added
- Add report display name, remove paths from stored files, fix sgRNA plot, CRISPRessoPooled report HTML, add citation to report
v2.0.25 - 02/21/2019
Added
- Add inferring of guides
v2.0.24 - 02/13/2019
Changed
- Update docker, setup.py
v2.0.23 - 01/24/2019
Added
- Add manifest.in
v2.0.22 - 01/23/2019
Changed
- Change license location, license update
v2.0.21 - 01/22/2019
Changed
- Detangled root location dependency from params
v2.0.20b - 01/22/2019
Changed
- Prepare for bioconda integration
License
CRISPResso2 is made available for free to academic researchers under this limited license for non-commercial use.
IMPORTANT: If you plan to use the CRISPResso2 for-profit, you will need to purchase a license. Please contact licensing@edilytics.com for more information.
CRISPResso2 END USER LICENSE AGREEMENT
BEFORE PROCEEDING, PLEASE READ THE END USER LICENSE AGREEMENT BELOW.
BY USING THIS SOFTWARE TOOL YOU ATTEST TO (I) BEING AN ACADEMIC RESEARCHER, (II) USING IT SOLELY FOR RESEARCH PURPOSES AND (III) YOUR ACCEPTANCE OF THE END USER LICENSE AGREEMENT.
-
General. As used herein, the term “you” or “your” means any individual or entity accessing this site or using the software tool “CRISPResso2” (the “Software Tool”) pursuant to this End-User License Agreement (“EULA”).
-
License to Use. The Software Tool is free for your use subject to the terms and conditions set forth below. The General Hospital Corporation, dba Massachusetts General Hospital (“MGH”) reserves the right to change, from time to time and at its sole discretion, this EULA. Your continued use of the Software Tool after any such modification constitutes your agreement and acceptance of such changes.
MGH owns all right, title and interest in the Software Tool. MGH grants to you, the “Licensee,” a royalty-free, non-exclusive, non-transferable, revocable license to use the Software Tool for non-commercial research or academic purposes only; it is NOT made available here as a free tool or download for any commercial or clinical use. You may not copy or distribute the Software Tool in any form. This license is limited to the individual that accesses the Software Tool. No right to sublicense or assign this EULA is granted herein.
The Software Tool optionally makes calls to unmodified versions of fastp https://github.com/OpenGene/fastp software, which is covered under its own license (MIT).
By using this Software Tool, you agree to allow MGH the right to collect data and statistics (i) on system usage patterns and (ii) to improve this Software Tool.
-
Limitations on Use. THE SOFTWARE TOOL HAS NOT BEEN REGISTERED OR APPROVED BY THE U.S. FOOD AND DRUG AGENCY, OR ANY OTHER GOVERNMENTAL AGENCY. THE SOFTWARE TOOL MAY BE USED ONLY AS A REFERENCE TOOL AND FOR CLINICAL EDUCATION, SIMILAR TO THE USE OF A TEXTBOOK OR A JOURNAL ARTICLE. THE SOFTWARE TOOL SHALL NOT BE USED AS A DIAGNOSTIC DECISION MAKING SYSTEM AND MUST NOT BE USED TO MAKE A CLINICAL DIAGNOSIS OR REPLACE OR OVERRULE A LICENSED HEALTH CARE PROFESSIONAL'S JUDGMENT OR CLINICAL DIAGNOSIS.
-
Disclaimer of Warranties. TO THE FULLEST EXTENT PERMITTED BY LAW, MGH PROVIDES THE SOFTWARE TOOL "AS IS" AND “AS AVAILABLE” WITH ALL FAULTS, ERRORS AND DEFECTS, AND NEITHER MGH NOR ANY OF ITS PERSONNEL NOR ANY OF ITS AFFILIATES IS RESPONSIBLE FOR ENSURING THAT ANY USE OF SOFTWARE TOOL WILL BE CLINICALLY SOUND, WITHOUT ERROR, UNINTERRUPTED OR OTHERWISE SUCCESSFUL. THE RIGHTS GRANTED IN THIS EULA ARE MADE AVAILABLE WITHOUT WARRANTY OF ANY KIND, EITHER EXPRESSED OR IMPLIED, INCLUDING, BUT NOT LIMITED TO, IMPLIED WARRANTIES OF MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE, TITLE AND NON-INFRINGEMENT.
-
Limitation of Liability. TO THE FULLEST EXTENT PERMITTED BY LAW, MGH SHALL NOT BE LIABLE TO YOU FOR ANY INDIRECT, INCIDENTAL, SPECIAL OR CONSEQUENTIAL DAMAGES (INCLUDING, BUT WITHOUT LIMITATION, ANY DAMAGES RESULTING FROM LOSS OF USE OR LOST BUSINESS, REVENUE, PROFITS, DATA OR GOODWILL) ARISING IN CONNECTION WITH YOUR USE OF THE SOFTWARE TOOL OR OTHERWISE, WHETHER IN AN ACTION IN CONTRACT, TORT, STRICT LIABILITY, NEGLIGENCE OR OTHERWISE, EVEN IF ADVISED OF THE POSSIBILITY OF SUCH DAMAGES.
-
Indemnification. You agree to defend, indemnify and hold harmless MGH and its affiliates, trustees, officers, employees, staff members, agents or contractors from and against any claim, charge, demand, action or suit, whether in contract, tort, strict liability, negligence or otherwise, for any and all losses, costs, charges, claims, demands, fees, expenses or damages of any nature or kind arising out of, connected with or resulting from (i) the use of the Software Tool by you, your affiliates, employees, staff, faculty, students, agents or (ii) relating in any way to this EULA.
In consideration of MGH providing access to the Software Tool free of charge, you agree not to bring any claim, lawsuit, or action (“Claim”) for any damages, costs, liabilities, settlement amounts and/or expenses (including attorneys’ fees) against MGH or its affiliates, trustees, officers, employees, staff members, agents or contractors arising out of or related to your use of the Software Tool.
- No Other Rights. You do not have the right to use the name, trademark, service mark, logo or other identifying characteristics of MGH or any of its affiliates or employees. All rights not expressly granted herein are reserved by MGH.
MGH may terminate your access to and use of the Software Tool at any time, with or without notice, for any reason or for no reason at all.
-
Governing Law. The construction and performance of this EULA will be governed by the laws of the Commonwealth of Massachusetts, without regard to conflicts of laws principles.
-
Entire Agreement. This EULA sets forth all of the covenants, provisions, agreements, conditions, and understandings between the parties regarding the subject matter herein, and there are no covenants, promises, agreements, conditions, or understandings, either oral or written, between them other than those set forth herein.
Should you have any concerns regarding this EULA contact us at licensing@edilytics.com.
Cite
For more on how CRISPResso works read the freely available published paper here.
If you like CRISPResso please support us by citing it in your work:
Clement K, Rees H, Canver MC, Gehrke JM, Farouni R, Hsu JY, Cole MA, Liu DR, Joung JK, Bauer DE, Pinello L.
CRISPResso2 provides accurate and rapid genome editing sequence analysis.
Nat Biotechnol. 2019 Mar; 37(3):224-226. doi: 10.1038/s41587-019-0032-3. PubMed PMID: 30809026.
@article{clement2019crispresso2,
title={CRISPResso2 provides accurate and rapid genome editing sequence analysis},
author={Clement, Kendell and Rees, Holly and Canver, Matthew C and Gehrke, Jason M and Farouni, Rick and Hsu, Jonathan Y and Cole, Mitchel A and Liu, David R and Joung, J Keith and Bauer, Daniel E and others},
journal={Nature biotechnology},
volume={37},
number={3},
pages={224--226},
year={2019},
publisher={Nature Publishing Group US New York}
}
Pinello L, Canver MC, Hoban MD, Orkin SH, Kohn DB, Bauer DE, Yuan GC.
Analyzing CRISPR genome-editing experiments with CRISPResso.
Nature biotechnology. 2016 Jul;34(7):695-7.
@article{pinello2016analyzing,
title={Analyzing CRISPR genome-editing experiments with CRISPResso},
author={Pinello, Luca and Canver, Matthew C and Hoban, Megan D and Orkin, Stuart H and Kohn, Donald B and Bauer, Daniel E and Yuan, Guo-Cheng},
journal={Nature biotechnology},
volume={34},
number={7},
pages={695--697},
year={2016},
publisher={Nature Publishing Group US New York}
}
CRISPResso Documentation
Select a version below: